International benchmarking study confirms APAF’s superior performance using SWATH mass spectrometry for proteomics
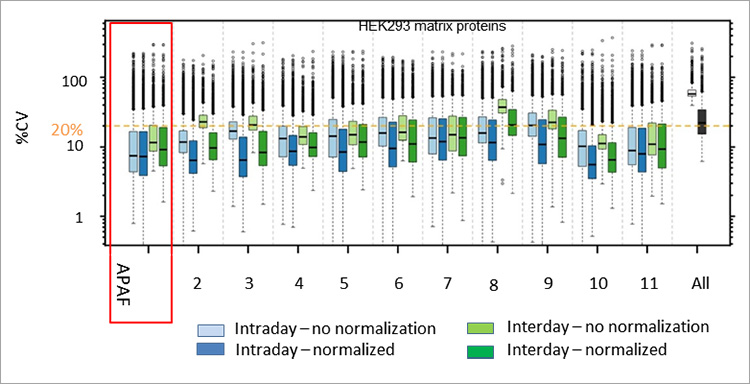
In a study published in Nature Communications, a multi-laboratory international assessment of SWATH mass spectrometry (also known as data-independent acquisition, or DIA) confirms excellent reproducibility and quantitative performance this this technique for proteomics. The Australian Proteome Analysis Facility (APAF) was selected as the only Australian participant in this study of 11 international laboratories and confirms APAF’s superiority to reliably detect and quantitate peptides from complex human cell lysates using SWATH-MS. APAF’s median intra-day and inter-day CV for quantitating 4077 proteins was ~7% and ~9%, respectively, values that were lower than the medians reported for the entire 11 laboratories.
The study reported the proteome quantitation of >4000 proteins from >200 experiments across all 11 participating labs, a feat described as unprecedented and one that could not be performed to such a high level of completeness using a conventional data-dependent MS sampling approach. This is an important study that provides broad confidence in the SWATH-MS approach to examine large studies of many samples such as cohort clinical studies or large-scale perturbation screens.
APAF has been providing SWATH-MS analysis to Australian researchers since 2013 and has recently described the bioinformatics tool SwathXtend to assist in reference library generation and analysis.
Collins BC, Hunter CL, Liu Y, Schilling B, Rosenberger G, Bader SL, Chan DW, Gibson BW, Gingras A-C, Held JM, Hirayama-Kurogi M, Hou G, Krisp C, Larsen B, Lin L, Liu S, Molloy MP, Moritz RL, Ohtsuki S, Schlapbach R, Selevsek N, Thomas SN, Tzeng S-C, Zhang H, Aebersold R. Multi-laboratory assessment of reproducibility, qualitative and quantitative performance of SWATH-mass spectrometry. Nat. Commun. 2017, DOI: 10.1038/s41467-017-00249-5.
Wu JX, Song X, Pascovici D, Zaw T, Care N, Krisp C, Molloy MP. SWATH mass spectrometry performance using extended peptide MS/MS assay libraries. Mol. Cell. Proteomics. 2016, 15(7), 2501-14.